Applying the AI: Lifting the Molecular Brakes on Nerve Cell Regeneration
Atomwise scientists are using machine learning to help University of Alberta researchers find compounds that have the potential to regrow damaged nerves.
Nerve damage is a common problem following injury in neurological disorders, but regrowing damaged nerves is difficult — especially if the underlying condition has been around for a long time. Even if the underlying disease is curable, nerve damage can still persist.
In the Neuroscience and Mental Health Institute at the University of Alberta, scientists in the laboratory of Douglas Zochodne have been trying to figure out new strategies that can speed up the nerve regeneration process by temporarily lifting the brakes on cell growth. Atomwise scientists are using the AtomNet® technologyto help the team find compounds to target proteins that prevent nerve regeneration. If the project succeeds, the compounds they identify could be used to treat nerve damage caused by trauma, diabetes, and other neuropathies that cause paralysis and sensory loss.
One target the scientists are interested in is retinoblastoma protein, Rb1. This protein is widely expressed in adult sensory neurons and their axons during regeneration, which makes it very appealing to scientists. Zochodne, a neurologist and the director of the university’s Neuroscience and Mental Health Institute, and his team had the idea that temporarily reducing Rb1 expression in nerve cells might be enough to trigger nerve regeneration.
Previously the scientists used short interfering RNA (siRNA) knockdown assays to reduce Rb1 protein expression in adult sensory neurons in order to increase new neurite growth. But siRNAs are difficult to work with and rapidly broken down by the body, so the team began exploring chemical options that could disrupt the interaction between Rb1 and a transcription factor called E2F1.
“Small molecules are a lot easier because you can make them go where you need them to in the body,” Zochodne explains. “Also, we only want to stop Rb1 temporarily, perhaps for a couple of weeks so the nerves can grow. Our idea is to find a drug that we can deliver locally in the nervous system for a short period of time.”
To identify optimal compounds for the task, the researchers needed to screen libraries containing millions of compounds. But doing this with real samples would have been cost-prohibitive. Instead, using computational tools to perform virtual screening would make the process quick and efficient. That led them to Atomwise.
“This team of subject matter experts had a lot of experience working with Rb1, which made this a very exciting collaboration for us,” says Tigran Abramyan, senior scientist at Atomwise.
Abramyan used the AtomNet® model to screen millions of commercially available compounds against Rb1. He searched for compounds using a general model trained on a curated dataset of proteins, their corresponding small-molecule ligands, and protein-ligand binding affinity information. He also applied theoretical filters to ensure that the models selected compounds capable of crossing the blood-brain barrier in the body.
“Protein-protein interactions are notoriously difficult targets for drug discovery in general and especially for virtual screens to make predictions about. The interfaces between partnering proteins are typically pretty shallow and hydrophilic, making it difficult for small molecules to bind to them,” Abramyan explains. But that was no problem for the AtomNet® model. “There is very little data available for Rb1 protein specifically. So, being able to find compounds using our general model that bind to Rb1 and disrupt Rb1-E2F1 association in vitro was a nice validation of our approach.”
The top 90 molecules predicted in silico to bind to Rb1 were selected for testing. These compounds were sent to Zochodne’s collaborator, Gary Eitzen, a cell biology professor, at the University of Alberta, for experimental testing in binding, functional and phenotypic assays.
Eitzen used surface plasmon resonance binding assays, which measure changes in the Rb1 protein when it is bound to a target compound, as well as functional assays, which assess what happens to nerve cells when Rb1 activity is disrupted. The scientists tested the effect of the compounds in two cell types — adult primary neurons and PC12 cells that differentiate into neuronal cells when they are exposed to nerve growth factors.
Assay results showed that many of the compounds identified by the virtual screens successfully bound to the Rb1 protein and disrupted the Rb1-E2F1 interaction. Both cell types showed evidence of neurite outgrowth and axonal regeneration when Rb1 was disrupted, which made it clear that the compounds were actually binding to a part of the protein that was important for its activity. The tests also revealed another interesting fact about the PC12 cells: when the researchers removed nerve growth factors and just used the compounds in the assays, these cells still showed neurite outgrowth and regeneration.
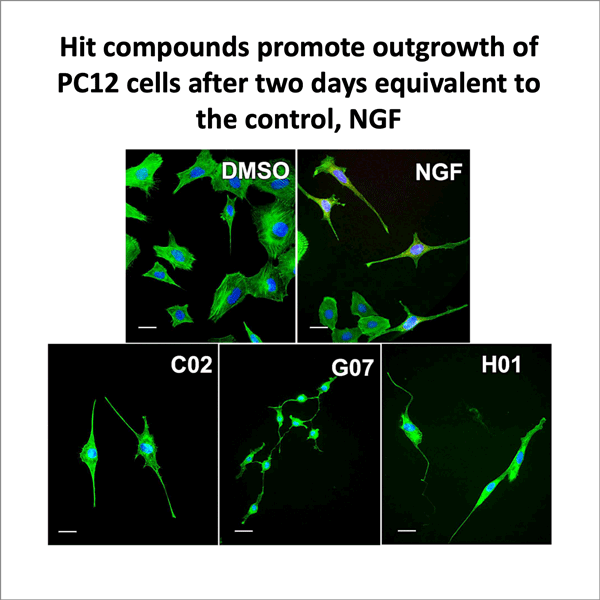
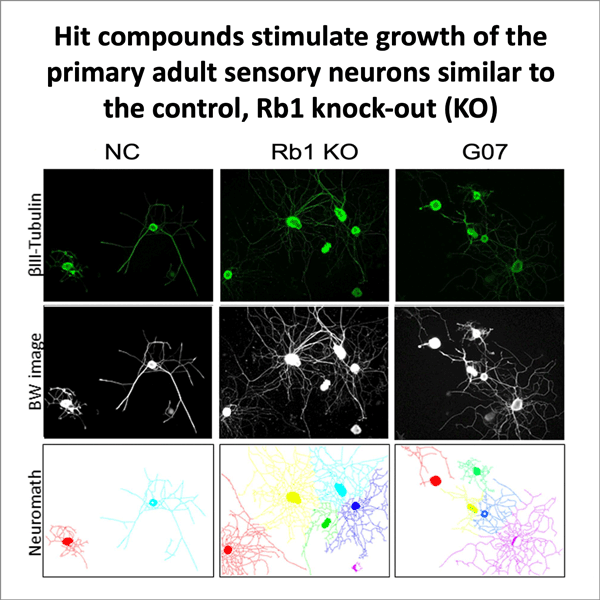
Images of assays showing PC12 cells (top image credit: Kendra Holte, Eitzen lab) and below, adult sensory neurons' response to hit compounds (bottom image credit: Prashanth Komirishetty and Aparna Areti, Zochodne lab)
The next step for this project is to perform hit expansion to identify more potent analogs of the compounds identified by the AtomNet® model. Eventually, the scientists will need to test the compounds in mice and rats to see if the nerve cells respond in vivo. “The cell assays that were used to test the compounds are useful predictors of how nerve cells will grow in rat or mouse models so they are a good starting point,” Zochodne says.
For Further Reading
Poster: Tigran Abramyan; Gary Eitzen; Douglas Zochodne, ACS 2022 Spring Meeting, Discovery of Small Molecule Ligands of Retinoblastoma Associated Protein Rb1 Using Artificial Intelligence.
About AIMS
The AIMS Award program, started in 2017, is designed to support promising researchers with resources that will help advance their work. AIMS Awards target research focused on finding solutions for complex human health conditions. To date, Atomwise has funded 7 rounds of AIMS Awards, completing over 100 collaborative projects and accepting over 800 projects into the program.
Related Posts
Subscribe
Stay up to date on new blog posts.
Atomwise needs the contact information you provide to send you updates. You may unsubscribe from these communications at any time. For information please review our Privacy Policy.
