Tackling TB: How Florida Scientists Are Working Toward a New Treatment
With as many as 2 million people dying each year from tuberculosis, there’s a crying need for better treatment options. Current therapy regimens involve a long and complicated combination of several drugs that many patients never complete, leading to the rise of drug-resistant TB strains.
What we need is a wonder drug, but what we have is a wonder bug. Major pharmaceutical companies have flooded resources into discovery and development programs for effective new treatments, but the pathogen responsible for TB is notorious for its ability to deflect or evade even the most pernicious compounds. By now, most biopharmas have completely abandoned efforts to conquer the disease.
 But Kyle Rohde is undeterred. An associate professor at the University of Central Florida, he is perhaps an unlikely protagonist in the therapeutic battle against TB. He has never worked in big pharma. He doesn’t have access to the high-throughput compound screening tools needed for traditional drug discovery. And his lab budget does not reach the millions of dollars usually required to identify, source, and test a large-scale compound library.
But Kyle Rohde is undeterred. An associate professor at the University of Central Florida, he is perhaps an unlikely protagonist in the therapeutic battle against TB. He has never worked in big pharma. He doesn’t have access to the high-throughput compound screening tools needed for traditional drug discovery. And his lab budget does not reach the millions of dollars usually required to identify, source, and test a large-scale compound library.
Still, Rohde has made progress where so many others have not. His motivation for persevering stems in part from a deep awareness of the situation: most pharmaceutical companies are not prioritizing new antibiotics, leaving the earlier-stage discovery work to academic scientists. “That puts us in a tough spot,” he says. “We don’t generally have the robotics or personnel or access to the huge drug libraries to test.”
Serendipity led Rohde to Atomwise, where he was selected for the company’s Artificial Intelligence Molecular Screen (AIMS) Award, a program that provides academic scientists with free access to customized virtual screening of millions or even billions of small molecules through the company’s AI-powered AtomNet® technology. To move the research forward even more, Atomwise sends dozens of small molecule compounds to each winner for follow-up functional testing.
For scientists in academia, drug discovery “becomes much more feasible with Atomwise’s virtual screens,” Rohde says. “You don’t have to randomly throw a million drugs on your bug and hope something kills it.” Instead of testing millions of compounds in real chemical screens, they’re tested in silico instead, with the AI model predicting how those compounds will interact with the target so scientists can focus only on the best candidates for follow-up in vitro testing. “Now it’s a size I can handle. It would be very difficult for me to screen a million compounds, but we could screen hundreds of compounds,” he adds.
In this AIMS Award project, Rohde submitted a list of 10 possible targets to the experts at Atomwise, who reviewed them and chose 2 they believed were most likely to succeed. After virtually screening millions of compounds for each, the Atomwise team, led by CADD Scientist Ha Truong, supplied Rohde’s team with 94 candidates for each target. His lab then performed bacterial assays to determine whether any of the compounds killed the TB pathogen. While several hits appeared promising, one compound stood out for its potency against the bacteria. Rohde’s team actually had to rerun the assay at lower concentrations because the first dosing series killed everything in its path. “It’s a lot more potent than we would have expected,” he says. Next, his lab will run biochemical assays to confirm that the compounds are acting on the intended target.
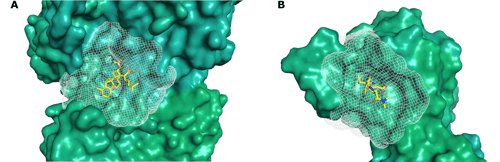 Models of known inhibitors bound to target site utilized in defining binding pockets for virtual screening. (A) Fusidic acid-bound pocket of T. thermophilus FusA, the template used to build homology model for Mtb FusA1. (B) Meropenem-bound pocket of Mtb LdtMT2.
Models of known inhibitors bound to target site utilized in defining binding pockets for virtual screening. (A) Fusidic acid-bound pocket of T. thermophilus FusA, the template used to build homology model for Mtb FusA1. (B) Meropenem-bound pocket of Mtb LdtMT2.
While traditional drug discovery scientists may wonder about the value of an AI-based screening process, Rohde is convinced. “Atomwise's strategy of leveraging neural networks and machine learning approaches has a proven track record,” he says. “They are also taking the data from all of these collaborations and feeding it back into the algorithm to make it even better.”
Rohde knows the road toward a new therapeutic will be a long one, but Atomwise has helped him achieve progress that was previously unthinkable for labs like his. The project has been approved for a second phase of the AIMS Award, so his team will benefit from another virtual screen intended to find analogs with similar structures that may have better pharmacological properties, as well as receive a second batch of compounds to test in functional assays. In addition, Rohde has submitted an NIH grant application in collaboration with Atomwise to accelerate his efforts — an application that would not have been possible without the assay results from compounds he got through the AIMS Award. “That’s really invaluable to put us in a position to compete for funding,” Rohde says.
The initial hits from his AIMS Award may or may not lead to a new treatment, but for now, Rohde is taking the win. “It’s still in the early stages, but I’m excited,” he says.
 Kyle Rohde and research lab member
Kyle Rohde and research lab member
Learn More
To learn more about Rohde’s research program, view his and Atomwise CADD Scientist, Ha Truong’s poster presentation at the American Chemical Society Spring 2021 National Meeting: Acceleration of TB Drug Discovery by AI-Based Virtual Screening Against Underexploited Targets
Related Posts
Subscribe
Stay up to date on new blog posts.
Atomwise needs the contact information you provide to send you updates. You may unsubscribe from these communications at any time. For information please review our Privacy Policy.