When Tech and Biotech VCs agree: Platform Companies and AI Drug Discovery
- by Abraham Heifets, co-founder and CEO of Atomwise
originally posted on Medium
In their cogent and insightful discussion about the different perspectives of Tech and Biotech investors, Drs. Shaywitz and Gibson make the implicit assumption that AI will be considered unproven by Biotech investors until assets picked by machines make it through clinical trials.
But “not yet in the clinic” is too powerful an objection — that benchmark would exclude many of the most exciting and best-funded biotech platform companies. To me, the more interesting questions are “Why do these platform companies get funded?” and “What would AI need to demonstrate to become part of that class?”. Because many of the most interesting therapeutic targets have zero lead compounds and are considered undruggable, the value of AI can be — and has been! — demonstrated much earlier.
Someone should tell the public CRISPR companies that you can’t get funded with a platform
The clinic is a powerful conceptual and practical boundary, and so it is easy to assume that the IND divides drug discovery into an easy-and-cheap domain before and a difficult-and-expensive domain after. David Shaywitz wrote,“Often, little, if any, value is placed on earlier-stage assets or the platform itself”, and Chris Gibson concurs when speaking about progress that“absent proof of concept data in human patients, is discounted severely.”
On the one hand, it’s undeniable that (1) assets have less risk and more value the closer they are to approval, and (2) no AI-discovered drug has made it into the clinic (although Recursion and Benevolent have repurposed drugs to new indications). However, the asset-in-the-clinic test is empirically not the bar that Biotech VCs employ. If it were, Biotech VCs would exclude all preclinical biopharma startups, including some of the extremely-well-funded CRISPR-, protein degradation-, and RNA-based platform companies of recent times.
What explains their success despite their early stage of technical and scientific development?
Drugging the undruggable is always valuable
Shaywitz helps us answer this question when he quotes David Weitz of Takeda. Although Weitz was speaking specifically about AI, the analysis is more broadly applicable:
“The value [of technology] is demonstrated when its predictions are validated and the combined [technology] prediction/validation effort is catalytic. Do [the technology’s] predictions provide insights not otherwise attainable by other methods? Are the predictions actionable and have a reasonable probability of success to merit experimental follow-up? Taken as a whole, was a meaningful roadblock more efficiently overcome than traditional methods?”
The excitement around CRISPR, protein degradation, and RNA-targeting techniques is justified because these techniques offer us the chance to drug fundamentally new targets that were not otherwise attainable by other methods.
How large does that chance need to be for the technologies to be worth pursuing? If any of these new technologies worked on 20% of undruggable targets, would that be valuable? 80%? 1%? Whatever your personal estimate, simple morality compels us to work on diseases that have not yet been addressed, rather than delivering compounds against well-characterized targets and be consigned to suffer the “better than the Beatles” problem in perpetuity.
The future of drug discovery is in using new technologies to drug the undruggable.
166 days to breakthrough
Consider PIM3, a cancer target that we’re pursuing at X-37, our latest joint venture, among several other challenging targets, such as Factor XIIa (anticoagulation), ZAP70 (autoimmune diseases), and SHP-2 (cancer).
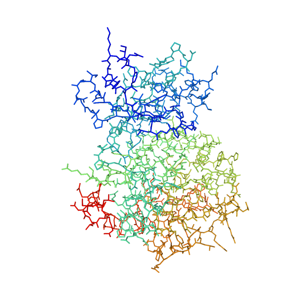
As with many kinases, the challenge with PIM3 is selectivity. PIM3 has been shown to be a good cancer target but the related protein PIM1 has associated cardiotoxicity.
Unfortunately, the ATP binding site of PIM3 differs in only 2 amino acids from the active site of PIM1. Furthermore, there is no publicly-known X-ray crystal structure for PIM3. In other words, the extreme similarity of the orthosteric sites, the absence of example selective molecules, and the lack of crystal structures make this target impossible for most computational techniques and challenging for human chemists.
Indeed, multiple pharma companies including Novartis, AstraZeneca, and AbbVie had worked on the PIM family and none had achieved the necessary selectivity for PIM3 over PIM1.
In contrast, we had discovered a selective inhibitor of PIM3 in 5.5 months — from the day we created the directory to initiate our analysis, to the day we received data from our CRO partner. To achieve this, Atomwise structural biologists proposed a novel allosteric site, built a detailed homology model, and screened massive purchasable libraries against it (we go up to 11B molecules now!).
Each of these steps is critical and each is a significant scientific and engineering effort. What makes this case study interesting is that, as soon as we find a selective compound, we have already demonstrated the ability to advance drug discovery beyond where traditional methods could take us empirically.
TL;DR
Targets where lead compounds are the limiting factor are hardly rare in drug discovery. Daniel Skovronsky, Chief Scientific Officer at Eli Lilly, said of the targets in the Atomwise collaboration, “Some of these are targets we’ve been working on for years without much progress. It’s tough to get a toe-hold, a place to start on a new target. If we can do that [through AI], we know how to take it the rest of the way.” In other words, when traditional drug discovery falls on the first hurdle, useful leads are real progress. Proof of utility happens very quickly because it does not take a clinical trial to prove that AI can help us drug the undruggable. AI has delivered exactly that — like CRISPR, protein degradation, and RNA-targeting.

Related Posts
Subscribe
Stay up to date on new blog posts.
Atomwise needs the contact information you provide to send you updates. You may unsubscribe from these communications at any time. For information please review our Privacy Policy.